High throughput quantitative PCR facility
Genomic Paris Centre – ENS PSL

Presentation
The qPCR-HD-GPC platform offers public and private research teams, as well as industry, expertise and methods for high-throughput quantitative genome and transcriptome analysis. The microfluidic technologies implemented on the platform enable all quantitative and digital PCR methods to be used for genomic analyses such as :
-
- Gene Expression variation (GE)
- Copy number variation (CNV)
- Search for polymorphisms (SNP)
Genotyping - Preparation and quantification of libraries for high-throughput sequencing
- Single cell genomics/trancriptomics
Absolute quantification (dPCR and cdPCR)
History
In September 2013, the qPCR-HD-GPC platform was opened to provide high-throughput quantitative PCR services available to the entire scientific community. The platform was then integrated into the Montagne Sainte-Geneviève technology platform network and offered new quantitative genomic analysis capabilities in connection with existing high-throughput sequencing services (ENS and Institut Curie), Laser Microdissection (Collège de France) and Flow Cytometry (ENS and Institut Curie).
Labels
The platform has been labelled IBiSA since 2018 (label renewed in 2021) and is part of the Genomic Paris Centre consortium, which includes the ENS Genomics and High Throughput qPCR platforms. Genomic Paris Centre is part of the core of the France Genomics Infrastructure since May 2020. In this sense, our objectives are to :
-
- Make quantitative genomic analysis technologies accessible to all laboratories
- Support research teams in the success of their projects
- Disseminate large-scale approaches in quantitative and functional genomics within the scientific community
- Train and inform on the advances in the field
years (since 2013)
related publications

projects
engineers
Who, what and how ?
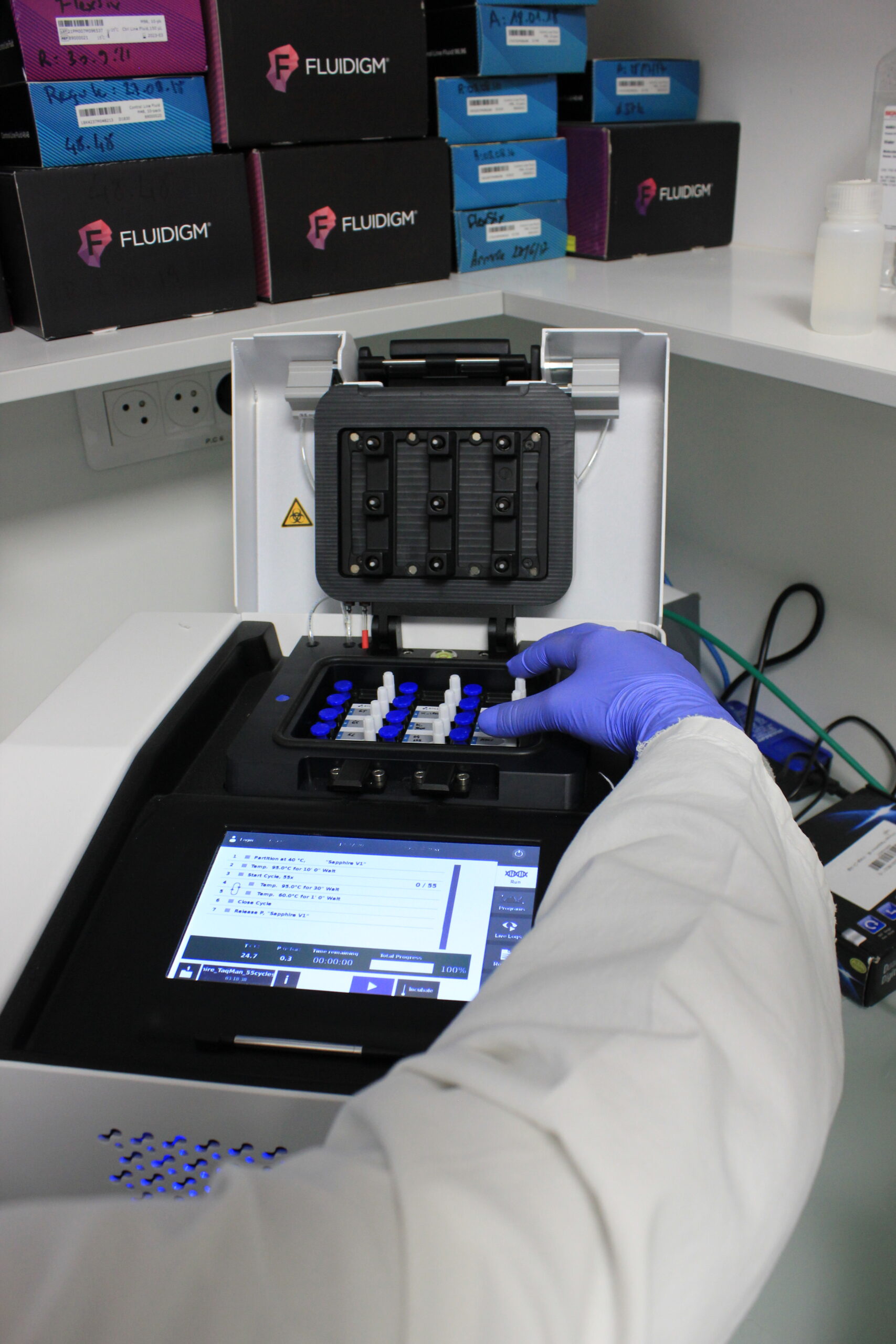
The qPCR-HD-GPC platform is handled by a biologist and a platform engineer. The platform is equipped with the Biomark-HDTM system from Fluidigm for quantitative and digital PCR in microfluidics, and with the NaicaTM system from Stilla Technologies for digital PCR in high-resolution nanodroplets (cdPCR). We offer a service based on a collaboration between the user and the qPCR-HD-GPC facility, that includes experimental design and analysis aimed at delivering publication-ready results. Supported samples include RNA, miRNA, cDNA, genomic DNA and environmental DNA from tissue or single cells. Quantitative and statistics analyses of the results are available with R.